Stewart Group
Our work focuses on two complementary aspects of genomics,
(I) mechanisms of epigenetic regulation in mammalian development and homeostasis;
(II) technologies of genetic engineering.
 © Neha Goveas
© Neha Goveas
Our research
Although the complete DNA sequence of an organism encodes the primary information, additional information is added by epigenetic regulation. In higher eukaryotes, epigenetic regulation is conveyed by covalent modifications of DNA (DNA methylation) and histone tails (mainly methylation, acetylation, ubiquitinylation).
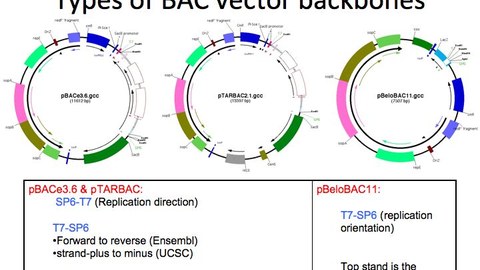
We have developed several genetic engineering innovations using site specific and homologous recombination, as well as transposition. Our aim is the fluent manipulation of mammalian genomes.