Bringmann Group
Why do we sleep? To answer this question, the Bringmann lab studies the regulation and functions of sleep in the roundworm C. elegans and in mice.
 © Yang Hu
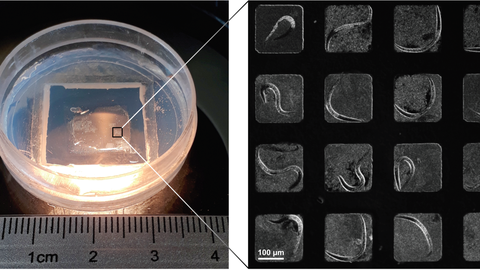
© Yang Hu
Our research
The molecular mechanisms underlying the vital functions of sleep in promoting health are not understood. This is surprising since sleep is essential for development and regeneration and sleep disorders are highly prevalent in industrialized societies, posing a massive unsolved medical and economic challenge.
The goal of our lab is to solve key sleep functions by analyzing genetic animal models of sleeplessness. The key model systems that we are studying are the nematode C. elegans as well as mice. Our goal is to obtain a molecular and mechanistic understanding of sleep functions in cellular health and well beeing. The results from animal models will thus provide a roadmap leading to the understanding and treatment of human sleep disorders and for developing regenerative therapies.